The battery compartments of the 26S Protein Recycling Machine
The degradation of proteins and the re-use of their basic building blocks is a process that is a matter of survival in cells. Researchers at the Max-Planck-Institute for Biochemistry present a detailed structure of the human protein recycling machine, the so-called 26S proteasome, in near-atomic resolution in their latest article published in PNAS. The high-resolution structure enabled the scientists to visualize the molecular energy carriers bound to the 26S proteasome, which provide the power for proteasome function. Detailed knowledge of the exact structure is the basis for the development of drugs for the treatment of cancers and neurodegenerative diseases.
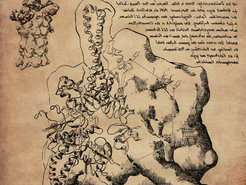
The sketch, drawn in the Da Vinci style shows the 26S proteasome structure, a part of the protein recycling system in human cells. The whole complex is shown in the left upper corner. Beneath, the component with the ‚battery compartments‘ (binding sites of ATP and ATP) is drawn in detail.
The visionary Leonardo da Vinci designed complex machines, such as helicopters, 500 years ago, which today are an integral part of our lives. To build these machines, he drew detailed construction plans. Today, structural biologists are reconstructing the frameworks of proteins, the molecular machines of our cells, at the highest resolution possible in order to precisely understand their function. However, these molecular machines are about 100 million times smaller than those da Vinci envisioned. To render these molecular-sized machines visible, special methods are required.
Scientists, along with Professor Wolfgang Baumeister and Dr. Friedrich Förster now show the detailed, three-dimensional structure of the 26S proteasome in nearly atomic resolution. The high-resolution structure of the 26S proteasome provides an abundance of knowledge about how the individual parts of the machine are connected to one another and how they work together. For the visualization of the structure, the scientists made use of cryo-electron microscopy.
In this method, the molecular machines isolated from cells are shock-frozen in thin films of water. As a result of this process, multiple copies of the cellular machines are present in various orientations. Thus, on images made through the electron microscope the proteasome can be seen from different angles. With the help of a computer, analogous to the computed tomography (CT) scan in medical imaging, a three-dimensional structure can be calculated.
Of special interest, the structure shows energy carrying molecules, adenosine phosphates, bound to the proteasome for the first time. “These molecules work in our cells like batteries. When a battery is charged, adenosine triphosphate (ATP) is present; if the energy is released, then adenosine phosphate (ADP) is created,” explains Andreas Schweitzer, first author of the study. “Now, we can study the ‘battery compartments’ of the proteasome precisely.” Why does the proteasome need energy? Prior to shredding the substrate (or target) protein into little pieces, the intricate three-dimensional structure of proteins is first ‘unfolded’. For this mechanical process, energy is required.
For that reason, the proteasome has 6 battery compartments, which are arranged in a hexagon. So far, it has not been known which of the battery compartments must be occupied at the same time so that the respective motor components may operate. With the help of the determined structure, the scientists were able to build an atomic model and show that in the idle state, i.e. at a point in time in which no protein is being degraded, all 6 compartments are filled with batteries. In this state, the energy of at least one of the batteries is already consumed; ADP is present, while the remaining compartments likely contain “charged” ATP molecules - significantly more than previously assumed.
“The current highest resolution structure of the 26S proteasome allows many new hypotheses about its function, which will help shape the research in this field. In the long term, this structure may serve as a basis for the development of medicines to treat specific types of cancer and neurodegenerative diseases,” theorizes Prof. Wolfgang Baumeister of the future. [AA]
Original Publication:
A. Schweitzer, A. Aufderheide, T. Rudack, F. Beck, G. Pfeifer, J. M. Plitzko, E. Sakata, K. Schulten, F. Foerster, and W. Baumeister: Structure of the human 26S proteasome at a resolution of 3.9 Å, PNAS, June 2016












