Andreas Bracher, PhD
Growth and survival of cells depends on correct protein folding and complex assembly. Molecular chaperones assist these tasks by recognizing folding intermediates, facilitating protein folding and prevention of protein aggregation. Moreover, molecular chaperones assist in the proteolytic removal of harmful mis-folded and aggregated species. Our work is aimed at the structure and function of molecular chaperones and associated proteins, using X-ray crystallography and cryo-electron microscopy as core techniques.
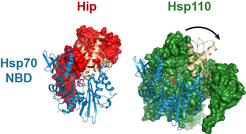
Specifically, we are interested in the regulation of Hsp70, which forms an important hub in the network of molecular chaperones in eukaryotic cells. We have determined structures of the transcription factor Hsf1, which upregulates expression of Hsp70 under protein-folding stress conditions. Hsp70 functions in protein folding by undergoing a nucleotide-dependent conformational cycle. In complex with ATP, protein substrate binding is highly dynamic while in the ADP and the nucleotide-free state, substrate is tightly bound. Co-chaperones regulate conformational cycling of Hsp70 either by triggering ATP hydrolysis or by exchange of ADP with ATP. For the latter reaction, several classes of nucleotide exchange factors (NEFs) have been identified in eukaryotes. Homologs to the prokaryotic NEF GrpE are found in mitochondria and chloroplasts. BAG domain proteins, Fes1p/HspBP1 and Hsp110 homologs are present in the cytosol and the endoplasmic reticulum (ER). Interestingly, these NEFs are only partially redundant and employ distinct mechanisms. Our crystal structures of the prototypical complexes of HspBP1 and of the S. cerevisiae Hsp110 protein Sse1p with Hsp70 revealed the details of their nucleotide exchange mechanisms. Moreover we solved the structure of the NEF antagonist Hip and showed how it stabilizes the Hsp70-ADP complex.
A paradigm for involvement of molecular chaperones in protein complex assembly is the biogenesis of the RuBisCO enzyme complex. The specific assembly chaperones RbcX, Raf1, Raf2 and Bsd2 found in cyanobacteria, green algae and plants assist in the biogenesis of this hexadecameric enzyme complex. Structures of Rubisco assembly intermediates determined in collaboration with the group of Dr. Manajit Hayer-Hartl revealed details of the assembly pathway.
A further interest is the structure-guided development of inhibitors for human FKBP51 in collaboration with Prof. Felix Hausch, University of Darmstadt, Germany.
People involved in the work
Leonie Mönkemeyer, Ph.D. student
