Proteomics identifies DNA repair toolbox
Max-Planck scientists identify protein profiles of DNA repair
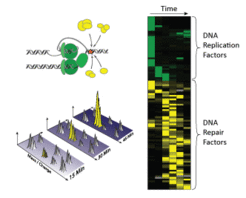
Identification of DNA repair proteins by mass spectrometry: Collision of the DNA replication machinery with lesions in the DNA triggers the recruitment of a large number of DNA repair factors (yellow) that help to repair the lesions. In proteomic analyses proteins are fragmented and identified by their mass and charge. Proteomic profiles (right) indicate the maximal accumulation of DNA replication and repair factors on the DNA.
During each cell division, more than 3.3 billion base pairs of genomic DNA have to be duplicated and segregated accurately to daughter cells. But what happens when the DNA template is damaged in such a way that the replication machinery gets stuck? To answer this question, scientists in the team of Matthias Mann at the MPI of Biochemistry, with colleagues in Copenhagen and at Harvard, have analyzed how the protein composition of the DNA replication machinery changes upon encountering damaged DNA.
To monitor such changes, they isolated DNA at several time points during the replication and repair process, and quantified the bound proteins using mass spectrometry based proteomics. In contrast to the conventional approach of quantifying a few selected proteins, proteomic analysis allows the simultaneous quantification of thousands of proteins from a biological sample. Using this unbiased approach, they found that as the replication machinery collides with lesions in the DNA more than 90 proteins are recruited to help repair the damaged DNA. These include many known DNA repair factors, as well as new proteins of previously unknown function. Together with the team of Professor Mailand at the Center for Protein Research in Copenhagen, Denmark, they are now analyzing whether and how these proteins promote the repair process. “Many of these recruited proteins are essential for the faithful propagation of genomic information to daughter cells,” explains Markus Räschle, first author of the study. “Because defects in DNA repair pathways are commonly observed in genetically inherited cancer syndromes, such as familially inherited breast cancer, it is possible that mutations in the new factors may contribute to genetic predisposition to cancers.”
DNA modifying agents are among the most frequently used class of chemotherapeutics. They work particularly well if the cancer cells they attack already have defects in the corresponding DNA repair pathways, as it frequently occurs in breast cancer and other tumors. Adaptation of proteomics for clinical diagnostics may open completely new possibilities. “Acquiring proteomic profiles directly from the tumor tissue may help oncologists to rapidly find the optimal class and dose of drugs,” says Matthias Mann, director at the Max Planck Institute of Biochemistry. “Our hope is that this approach may lead to more effective therapies with fewer side effects.”
Original publication:
M. Räschle, G. Smeenk, R.K. Hansen, T. Temu, Y. Oka, M.Y. Hein, N. Nagaraj, D.T. Long, J.C. Walter, K. Hofmann, Z. Storchova, J. Cox, S. Bekker-Jensen, N. Mailand, M. Mann: Proteomics reveals dynamic assembly of repair complexes during bypass of DNA crosslinks. Science, May 1, 2015
DOI: 10.1126/science.1253671
Abstract












