
Bioinformatics Core Facility
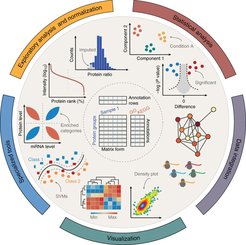
The aim of the Bioinformatics Core Facility is to assist wet-lab researchers in our institute for their bioinformatics data analysis. Our facility is tightly embedded in the Computational Systems Biochemistry and works in close contact with the researchers at the Max-Planck Institute of Biochemistry and Biological Intelligence. These close links ensure that our services remain state-of-the-art, not only by keeping up with novel developments in the field, but also by offering bioinformatics assistance that goes beyond publicly available analysis methods.
Our Bioinformatics core facility has several tasks to help the researchers at the Max-Planck Institute of Biochemistry and Neurobiology:
- Preliminary discussion about experimental design together with other core facilities, such as sequencing facility. This crucial step assures to obtain high quality results from your projects.
- Consultation for the bioinformatics methods, available computational tools and statistical data analysis
- Preparation of data sets to submit in public repositories, such as PRIDE, GEO or SRA, as requested by many journals upon manuscript submission
- Provide bioinformatics trainings
Some selected topics from our bioinformatics core facility:
Genomics
Next-Generation sequencing (NGS) data analysis
Comparative genomics
Combining chromatin immunoprecipitation assays with sequencing (ChIP-seq)
Whole genome sequencing (WGS)
Metabolomics
Proteomics
Shot-gun proteomics analysis (both identification and quantification)
De novo sequencing
Chemical crosslinking combined with mass spectrometry
Proteogenomics
Protein-protein interactions
Ion-mobility mass spectrometry data analysis
Multiple sequence alignment
Alternative splicing
Protein functional annotation and prediction
Pathway enrichment analysis
Phylogenetic analysis
Motif search
Sequence similarities (BLAST)
Transcriptomics
Single cell transcriptomics
RNA-seq for mRNA, or long non-coding RNA
tRNA-Seq
Ribosome profiling
RNAi screening
Single guide RNA for CRISPR
Multi-omics
We are planning to organize a one-day workshop on Computational Mass Spectrometry-based Proteomics that includes hands-on training on how to run MaxQuant and Persesus.
You can find some links for several useful links about Bioinformatics and programming.
| Protein structure visualization, prediction & analysis | PyMol, UCSF-Chimera, HADDOCK, I-TASSER |
| Genomics | |
|
Programming tutorials |
learn programming: R, R studio python and documentation for python 2.7, python 3.7.1and jupyter Johns Hopkins Data Science Lab: courses, books programming exercises: coding-bat for python or coding-bat for java |
| Annual MaxQuant summer school | the largest workshop in computational proteomics, details and how to register here |
| Proteomics analysis | MaxQuant, Perseus |
| Databases | GEO, DAVID, HumanProteinAtlas Uniprot, Ensembl |
| Public repositories | PRIDE, ProteomeXchange Consortium |
| Statistics | Bioconductor, Perseus |
